About plantit#
What is it?#
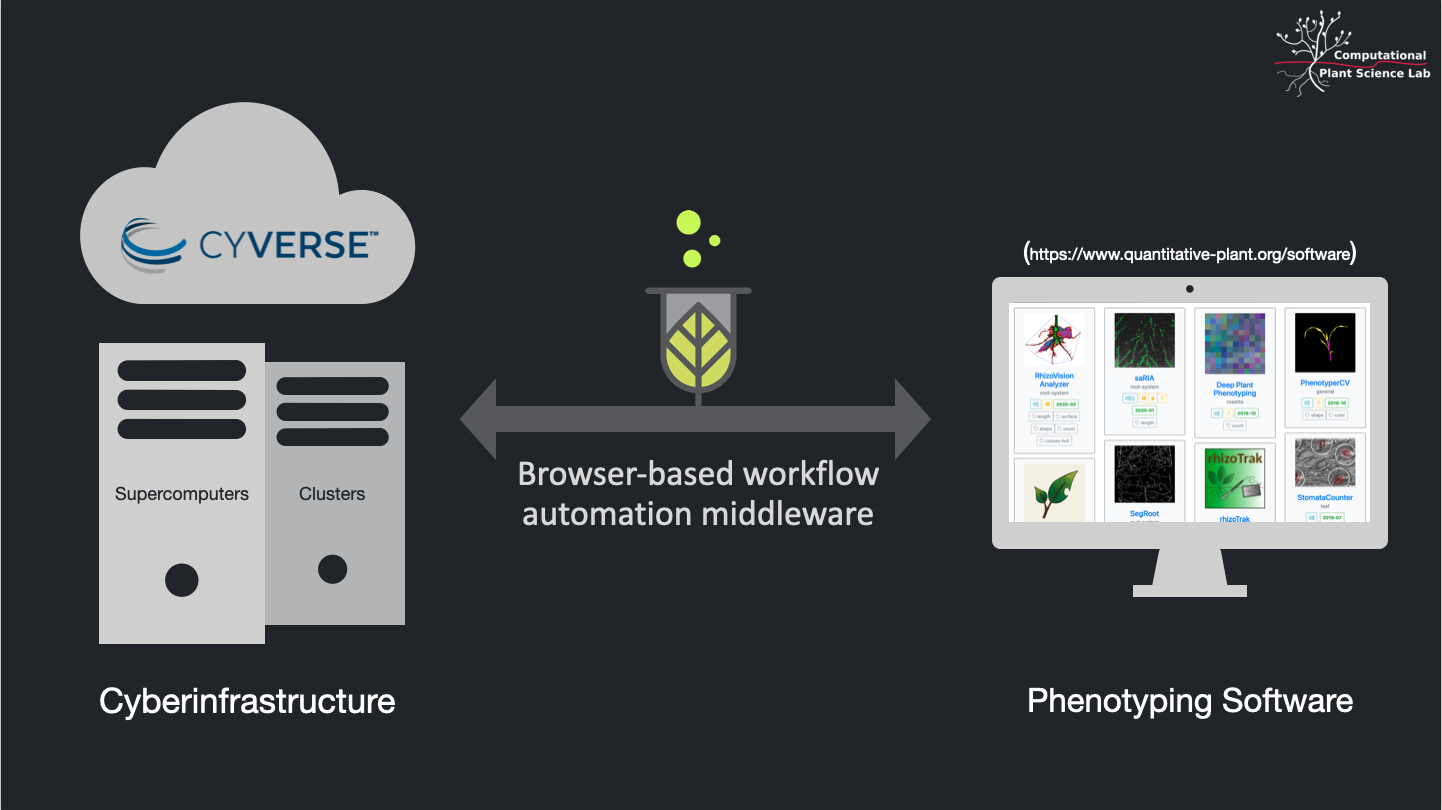
plantit is a workflow automation tool for computational plant science. It is simultaneously software-as-a-service for researchers and a platform-as-a-service for programmers & developers.
 Researchers
Researchers
SaaS for researchers: store, publish, and access data with CyVerse, run (possibly highly parallel) simulations and analyses from a browser.
 Developers
Developers
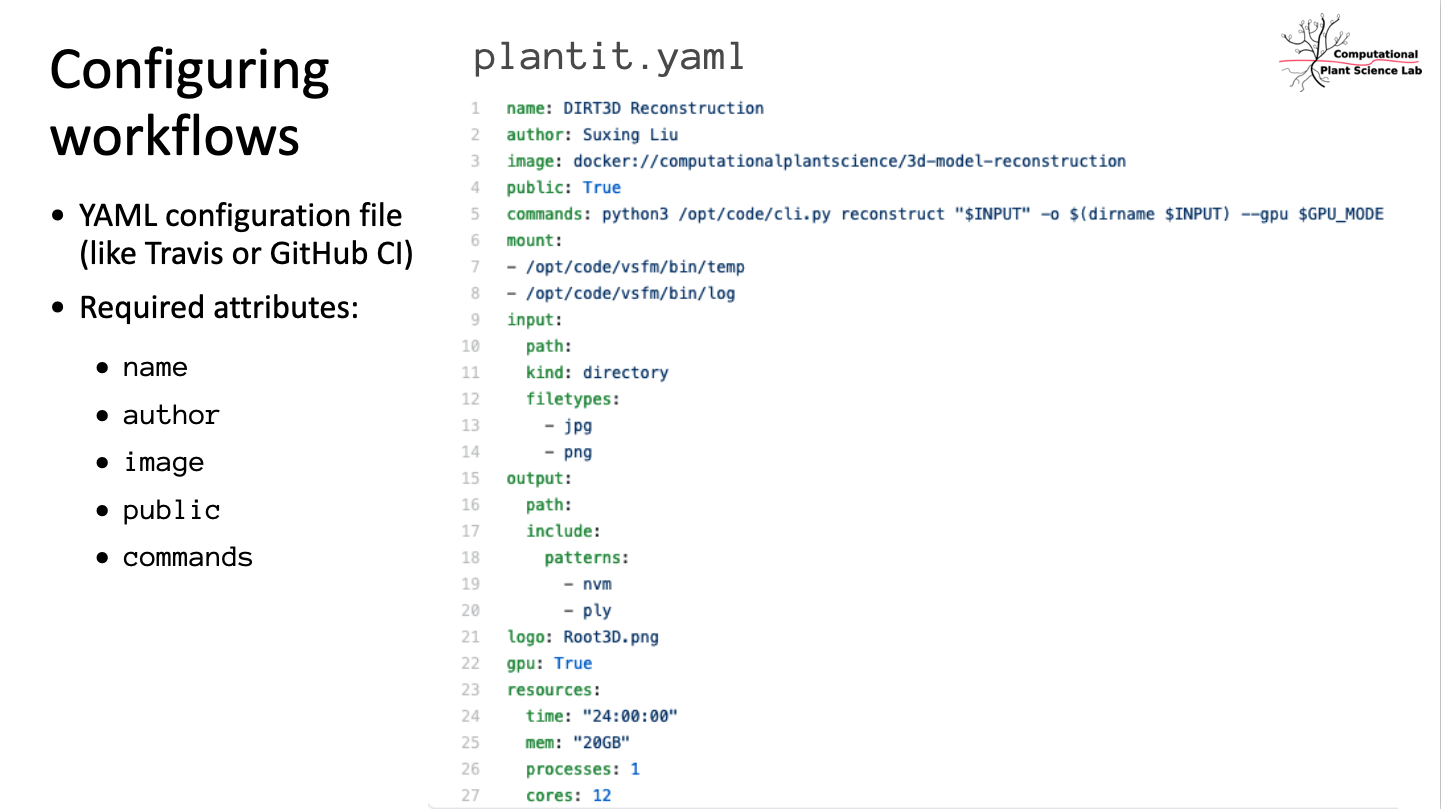
Paas for developers: built on GitHub and Docker, add a plantit.yaml file to any public GitHub repository to deploy Docker images as Singularity containers on clusters or supercomputers.
What isn’t it?#
plantit is none of the following (although it tries to glue these systems together in helpful ways).
a pipeline orchestrator (e.g., Snakemake, Nextflow, Luigi, Airflow, Metaflow)
a distributed queue or task scheduler (e.g., Celery or Dask)
a batch processing, streaming, or analytics platform (e.g., map-reduce or Spark)
a container automation system (e.g., Kubernetes)
a cluster scheduler (e.g., Torque/Moab, Slurm)
What can I use it for?#
plantit is flexible enough to run most container-friendly workloads. If your software can be packaged with Docker and invoked on the command line, plantit can probably run it. That said, plantit is designed primarily for batch processing images in various phenotyping contexts. If you want to do genomics, an established tool like CoGe or easyGWAS may be a better fit. Feel free to get in touch with questions about your use case.