Quickstart#
Use cases#
plantit aims to support two user groups with different concerns and priorities.
researchers: analyzing data, running models & simulations (submitting workflows)
developers: publishing and maintaining research software (publishing workflows)
With plantit the latter group can quickly and easily publish an algorithm to a broader, possibly non-technical user community. In this way it’s a continuous deployment tool. It’s also a science gateway, hosting plug-and-play algorithms in a web GUI such that any user can leverage XSEDE HPC/HTC clusters for high-throughput phenomics, no programming experience required.
Conceptual model#
plantit has a few fundamental abstractions:
Dataset
Agent
Workflow
Task
A Dataset is a set of data objects. A Workflow is a containerized research application. A workflow must yield a dataset as output and may accept one as input (workflows should be designed as functions or generators, not for their side effects — ideally, they should have none). An instantiation of a workflow is called a Task. An Agent is a cluster queue that can run tasks.
Datasets#
A Dataset is a collection of data objects in the CyVerse data store.
Agents#
An Agent is a deployment target: an abstraction of a cluster or supercomputer along with SLURM scheduler configuration details.
Workflows#
A Workflow is an executable research application packaged into a Docker image. Workflows are deployed in the Singularity container runtime. To define a workflow, add a plantit.yaml file to any public GitHub repository.
Tasks#
A Task is an instance of a workflow, deployed to an agent. When a task is submitted from the browser, the plantit web app hands it to an internal queue feeding an orchestrator process. When the orchestrator picks up the task, it generates a job script and submits it to the selected cluster/supercomputer scheduler, then monitors its progress until completion.
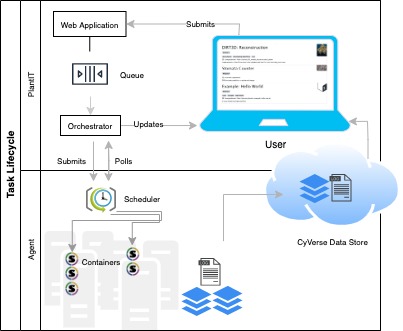 Interactions
Interactions
Projects#
A Project is a MIAPPE investigation, which may contain one or more studies. MIAPPE (Minimum Information About a Plant Phenotyping Experiment) is a formal ontology for organizing data, metadata, experiments, and analyses. plantit allows datasets and tasks to be freely associated with MIAPPE projects.
Submitting tasks#
Select a workflow#
To explore workflows, navigate to the Workflows tab from the home view.
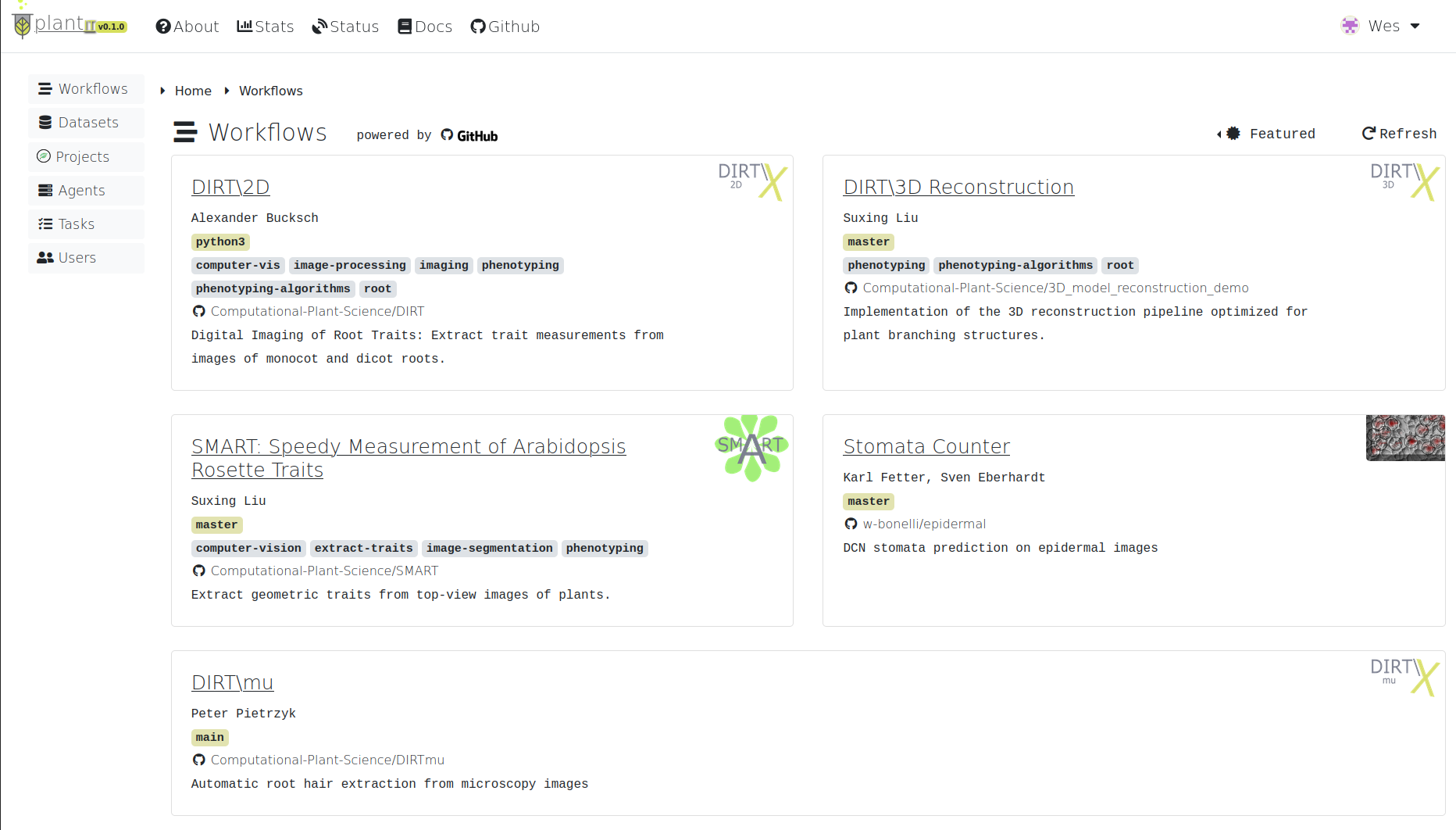 Workflows
Workflows
By default, this page will display the Featured workflow context: a curated set of applications provided by the Computational Plant Science lab, collaborators, and other researchers.
Click the Featured dropdown to select a different context. Options include:
Examples: a small set of simple workflows to serve as templates and examples
Public: all publicly available workflows
Yours: your own workflows (private and public)
[Organization]: workflows belonging to a particular organization
[Project]: workflows associated with a particular MIAPPE project
Select a workflow to view its authorship, related publications, parameter list, and deployment configuration.
Submit to an agent#
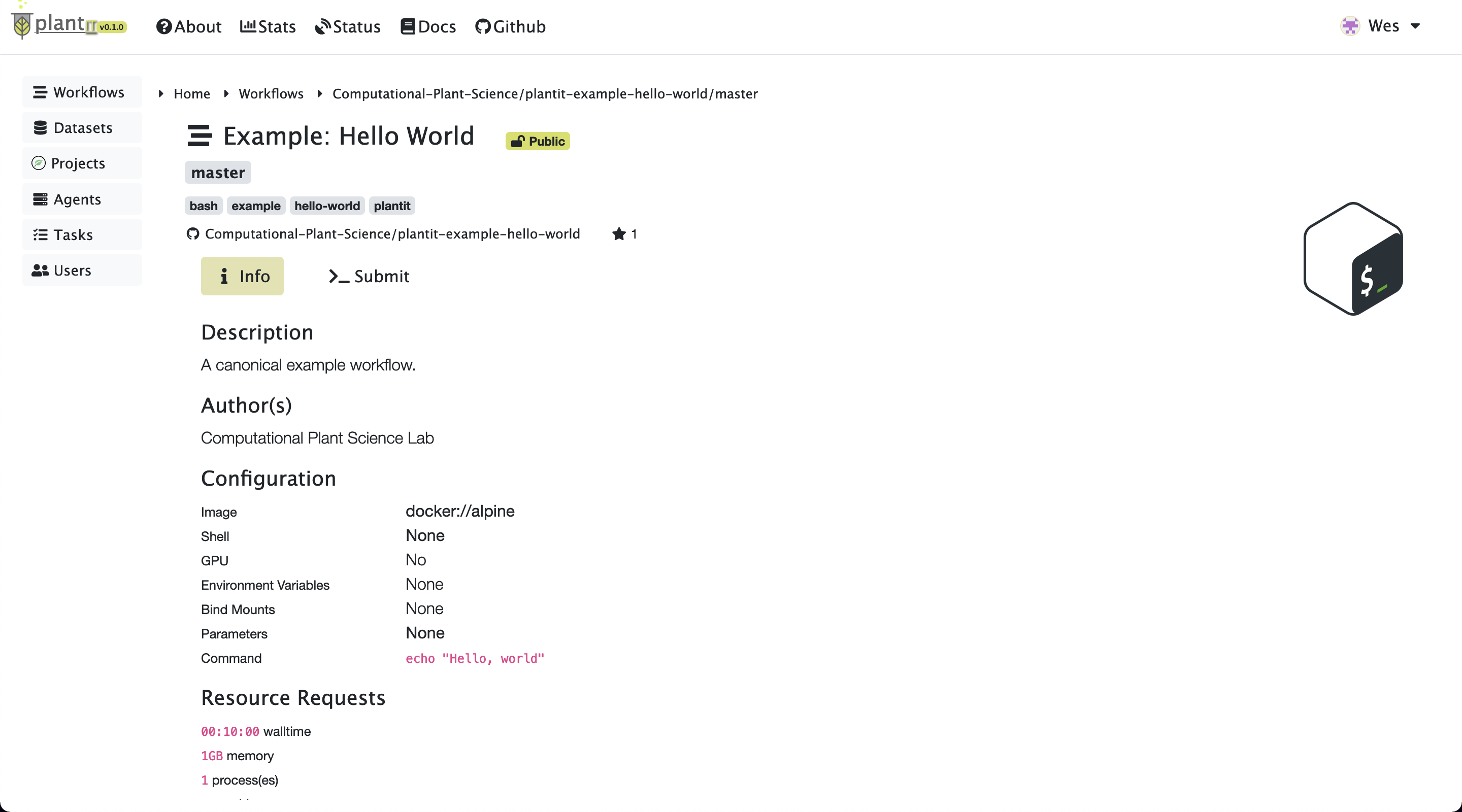 Task information
Task information
To configure and submit a task for the workflow you’ve selected, click Submit. This will present some configuration options including (at least):
ID: the task’s (unique) identifier
Tags: arbitrary text tags to associate with the task
Time: the task’s time limit
Agent: the agent to submit the task to
Output: the folder to deposit results in
If the workflow requires input files or parameters, corresponding configuration sections will be shown.
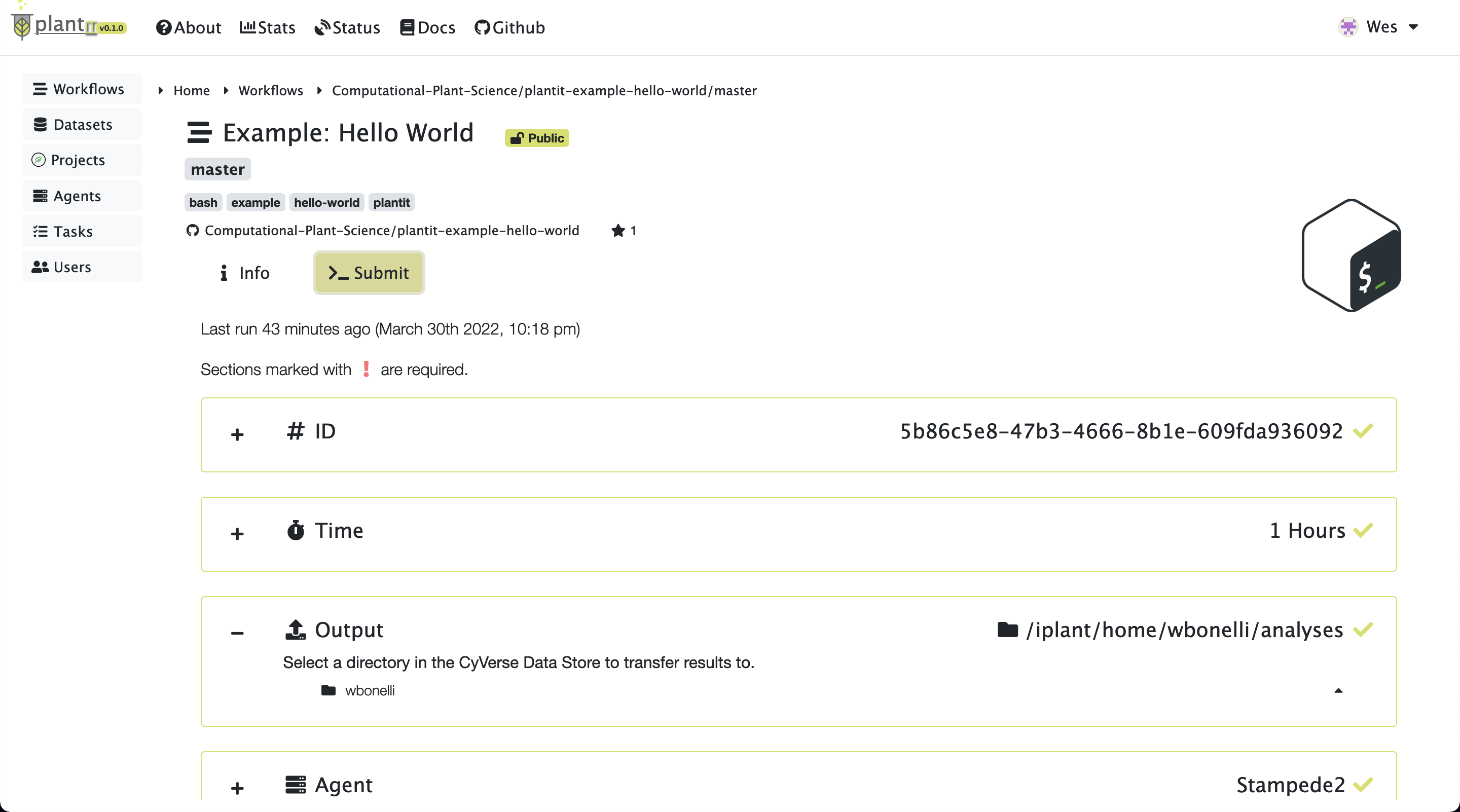 Task submission
Task submission
After all fields have been configured, click the Start button to submit the task.
Monitor status#
After a moment the task page will appear. At first there may be no log messages.
 Task status:
Task status: CREATED
Before long the task should be created, scheduled, and started on the appropriate agent. At this point you should see a few lines of log output:
 Task status:
Task status: RUNNING
When a task completes successfully, the status will change from RUNNING to COMPLETED.
 Task status:
Task status: COMPLETED
Retrieve results#
The output folder in the CyVerse data store section will eventually open at the bottom of the view (you may need to reload the page). Results will be zipped into a file with name matching the task’s ID.
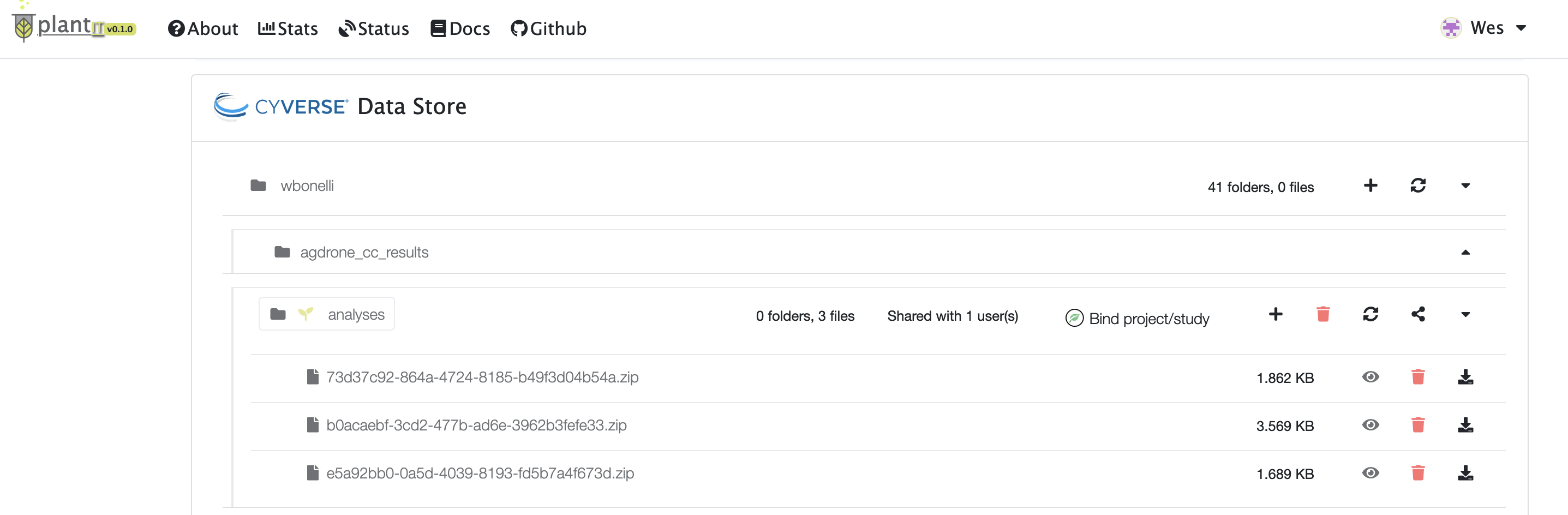 Task results
Task results